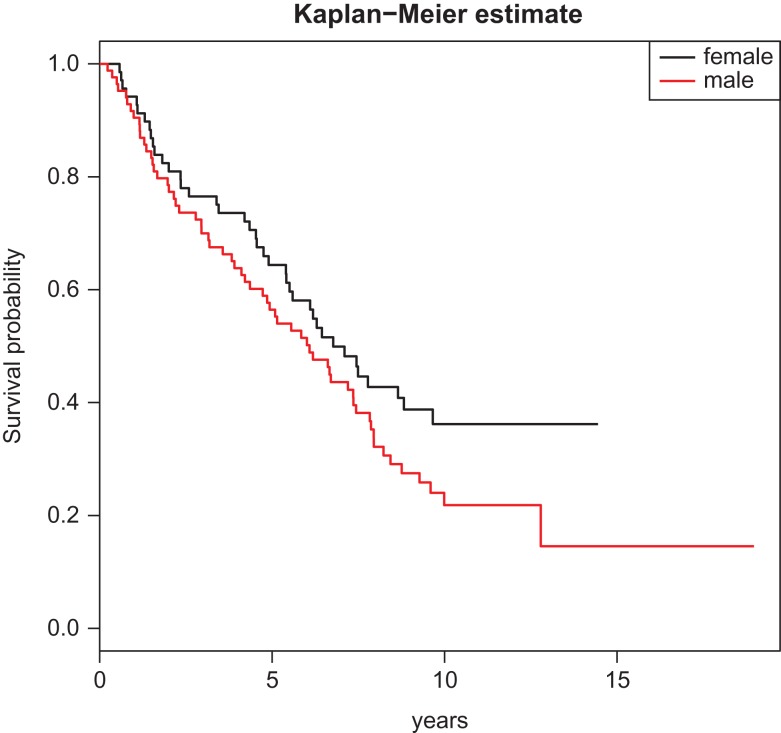
Quantile regression hyperlinks the entire distribution of an final result to the covariates of curiosity and has grow to be an vital various to generally used regression fashions.
However, the presence of censored data corresponding to survival time, typically the primary endpoint in cancer research, has hampered using quantile regression strategies due to the incompleteness of data.
With the appearance of the precision drugs period and availability of excessive throughput data, quantile regression with high-dimensional predictors has attracted a lot consideration and offered added perception in comparison with conventional regression approaches.
This paper offers a sensible information for utilizing quantile regression for proper censored final result data with covariates of low- or high-dimensionality.
We body our dialogue utilizing a dataset from the Boston Lung Cancer Survivor Cohort, a hospital-based potential cohort research, with the objectives of broadening the scope of cancer analysis, maximizing the utility of collected data, and providing helpful statistical options.
We use quantile regression to establish medical and molecular predictors, for instance CpG methylation websites, related to high-risk lung cancer sufferers, for instance these with brief survival.

Aligning the Aligners: Comparison of RNA Sequencing Data Alignment and Gene Expression Quantification Tools for Clinical Breast Cancer Research.
The speedy growth of transcriptomics and affordability of next-generation sequencing (NGS) applied sciences generate rocketing quantities of gene expression data throughout biology and drugs, together with cancer analysis.
Concomitantly, many bioinformatics tools have been developed to streamline gene expression and quantification. We examined the concordance of NGS RNA sequencing (RNA-seq) evaluation outcomes between two predominant packages for learn alignment, HISAT2, and STAR, and two hottest packages for quantifying gene expression in NGS experiments, edgeR and DESeq2, utilizing RNA-seq data from breast cancer development collection, which embody histologically confirmed regular, early neoplasia, ductal carcinoma in situ and infiltrating ductal carcinoma samples microdissected from formalin fastened, paraffin embedded (FFPE) breast tissue blocks.
We recognized vital variations in aligners’ efficiency: HISAT2 was liable to misalign reads to retrogene genomic loci, STAR generated extra exact alignments, particularly for early neoplasia samples. edgeR and DESeq2 produced comparable lists of differentially expressed genes, with edgeR producing extra conservative, although shorter, lists of genes.
Gene Ontology (GO) enrichment evaluation revealed no skewness in vital GO phrases recognized amongst differentially expressed genes by edgeR versus DESeq2.
As transcriptomics of FFPE samples turns into a vanguard of precision drugs, selection of bioinformatics tools turns into essential for medical analysis. Our outcomes point out that STAR and edgeR are well-suited tools for differential gene expression evaluation from FFPE samples.
