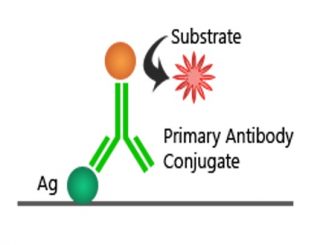
PCR OG Elisa Kit
Introduction The enzyme-linked immunosorbent assay (ELISA) is a labelled immunoassay that is considered the gold standard of immunoassays. This highly sensitive immunological test is used […]

Introduction The enzyme-linked immunosorbent assay (ELISA) is a labelled immunoassay that is considered the gold standard of immunoassays. This highly sensitive immunological test is used […]
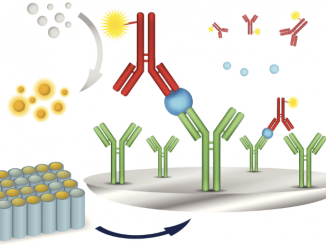
ELISA kits Optimized, high-quality Invitrogen ELISA kits allow you to measure target-specific proteins with confidence, reliability, and consistency. A variety of ELISA kit formats are […]
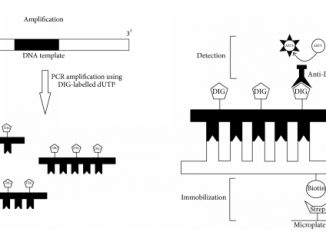
Abstract Enzyme-linked immunosorbent assay-polymerase chain reaction (PCR-i-ELISA) is an immunodetection method that can quantify the PCR product directly after immobilization of biotinylated DNA on a […]

Older patients expertise the next danger of treatment-related toxicity (TRT). The G8 screening tool was developed to separate cancer older patients match to obtain commonplace […]

The being pregnant fee after most cancers therapy for feminine survivors is decrease than that of the final inhabitants. Future infertility is a important concern […]

Body composition evaluation can present data related to breast most cancers sufferers’ (BCP) prognosis, that may lead interventions to enhance survival outcomes. The intention of […]
Going from bench to bedside is a simplified description of translational analysis, with the final word aim being to enhance the well being standing of mankind. Tumor […]
Cancer is among the most main causes of mortalities worldwide. It is brought on by the buildup of genetic and epigenetic alterations in 2 kinds […]
At the earliest contemplation of surgical procedure, the entire cohort was screened for impaired bodily, dietary, and/or psychological standing. Patients screened at increased danger acquired […]

FROZEN TISSUE PROCESSING TECHNIQUEThis article describes a tissue watering system for the preparation of frozen sections. The method differs from conventional target casting and is […]
Copyright © 2024 | WordPress Theme by MH Themes